Doc-Alignment
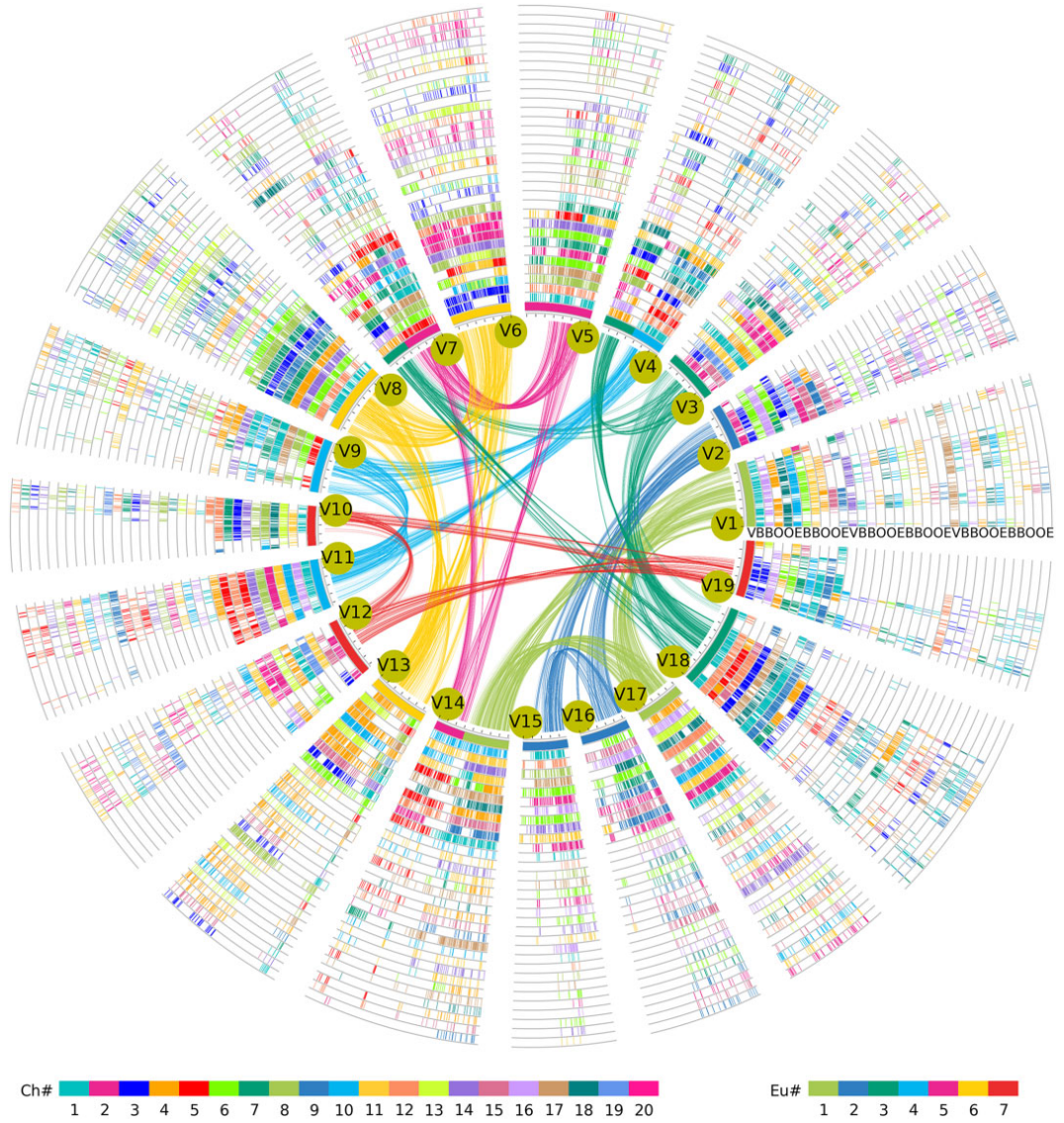
Inter-genomic and intra-genomic comparisons can help reveal the structural complexity of Cucurbitales genomes. We used V.vinifera as a reference, and by comparing homologous gene locus maps and Ks values between V.vinifera and other Cucurbitales, we could separate orthologous and paralogous genes produced by different polyploidization in the species genomes. We created a hierarchical lists of homologous genes using V.vinifera as a reference.
The relevant gene ids can be obtained from the Cucurbitales blast and match under the Tools module. This link is Cucurbitales blast and match.